RECENT PROJECTS
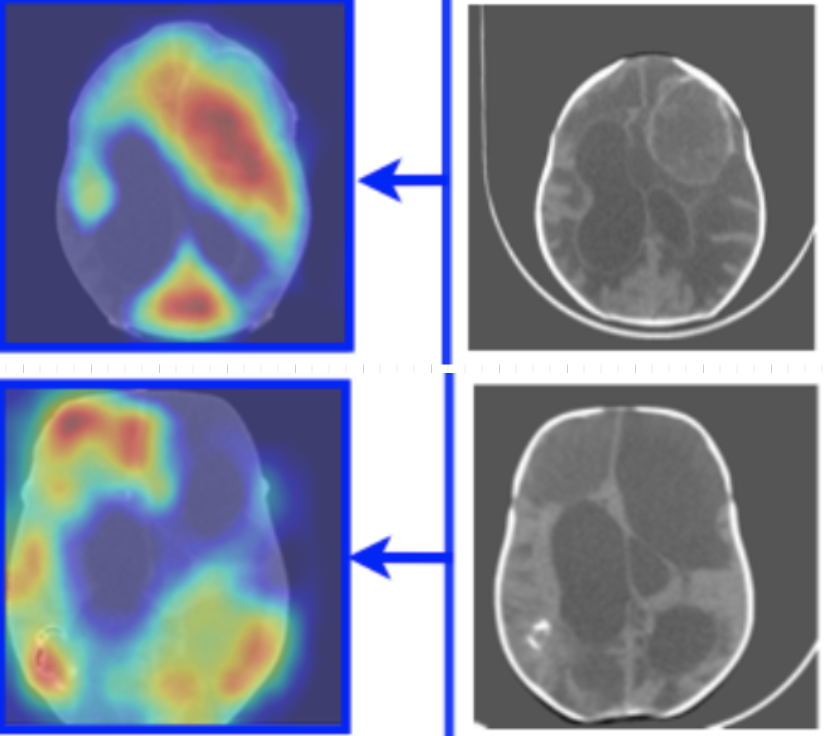
Infection Diagnosis of Hydrocephulus CT images: a domain-enriched attention learning approach
We develop two novel deep learning classification models with novel attention regularization for CT hydrocephalus classification and infection diagnosis (a 2D version and a 2D/3D hybrid version).
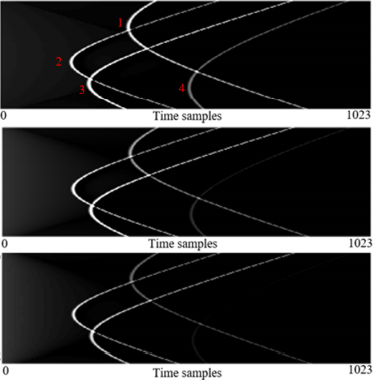
Simultaneous Denoising and Localization Network for Photoacoustic Target Localization
We develop a novel deep learning method designed to explicitly exhibit robustness to noise present in photoacoustic radio-frequency (RF) data.
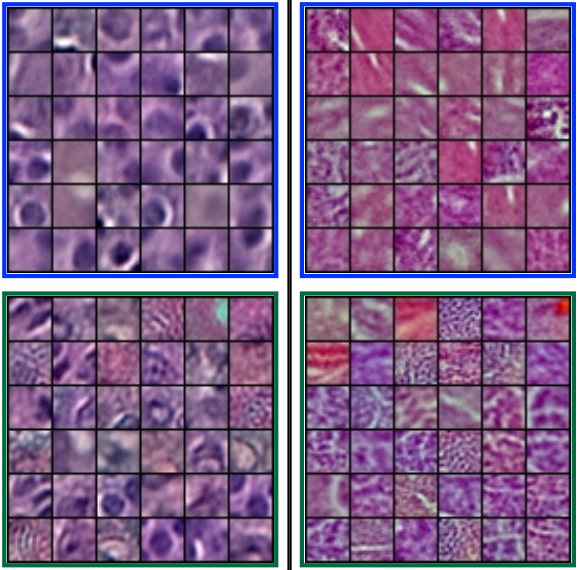
Histopathological Image Classification using Discriminative Feature-Oriented Dictionary Learning
We propose an automatic feature discovery framework for extracting discriminative class-specific features and present a low-complexity method for classification and disease grading in histopathology.
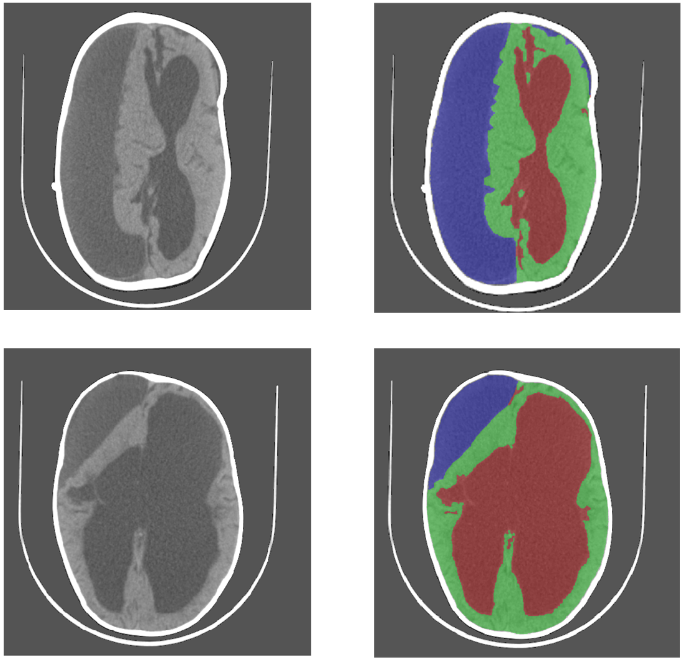
Learning Based Segmentation of CT Brain Images: Application to Post-Operative Hydrocephalic Scans
Most intensity and feature based segmentation methods fail to separate subdurals from brain and CSF as subdural geometry varies greatly across different patients and their intensity varies with time. We combat this problem by a learning approach that treats segmentation as supervised classification at the pixel level, i.e. a training set of CT scans with labeled pixel identities is employed.
Deep MR Image Super-Resolution Using Structural Priors
Unlike regular optical imagery, for MR image super-resolution generous training is often unavailable. We therefore propose the use of image priors, namely a low-rank structure and a sharpness prior to enhance deep MR image superresolution.
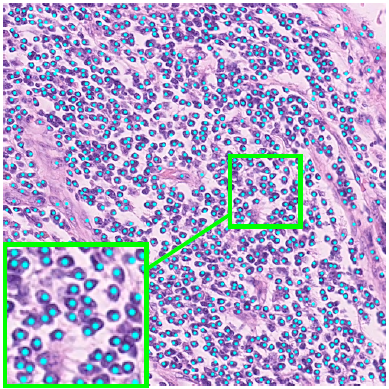
Deep Networks With Shape Priors For Nucleus Detection
Nuclei detection has been a topic of enduring interest with promising recent success shown by deep learning methods. These methods train for example convolutional neural networks (CNNs) with a training set of input images and known, labeled nuclei locations. Many of these methods are supplemented by spatial or morphological processing.
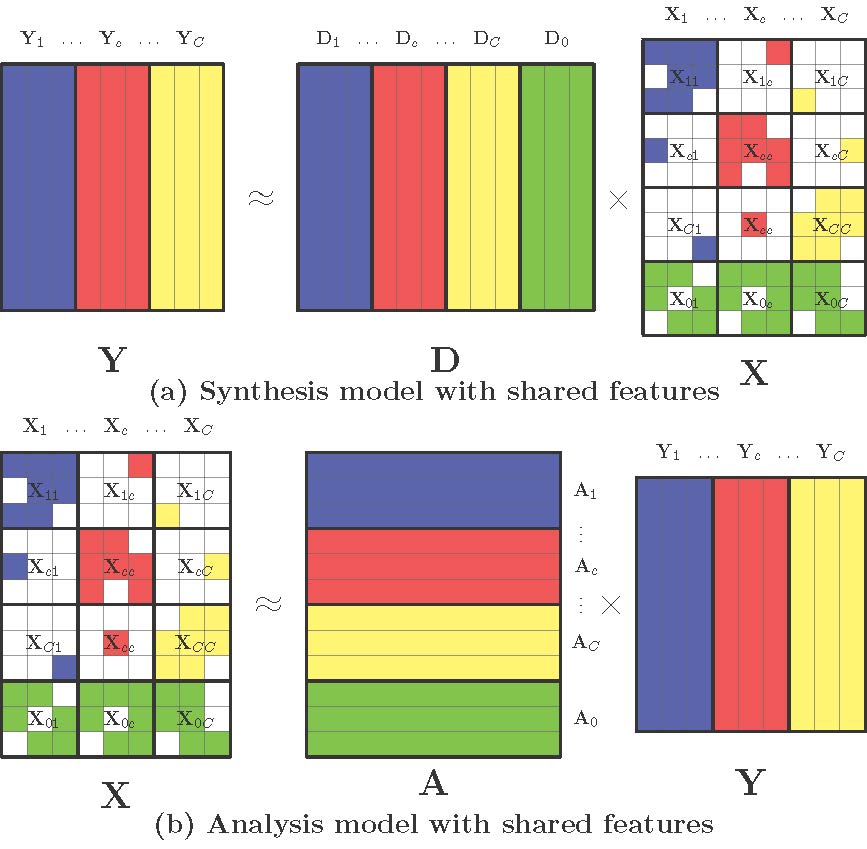
ALSF - Analysis-Synthesis learning with shared features for histopathological image classification
The diversity of tissue structure in histopathological images makes feature extraction for classification a challenging task. We introduce the learning of a low rank shared dictionary and a shared analysis operator, which more accurately represents both similarities and differences in histopathological images from distinct classes.
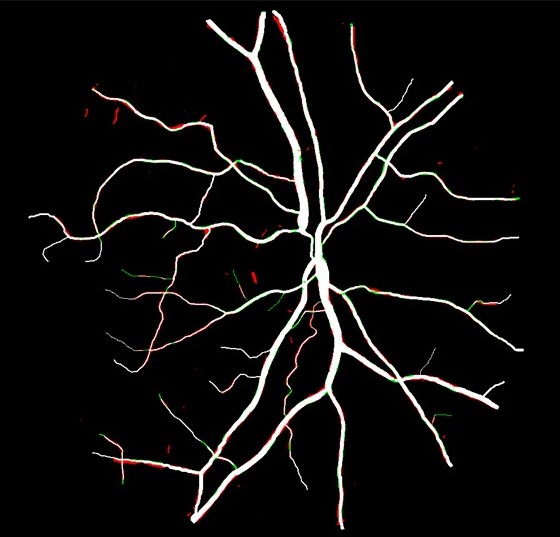
Deep Retinal Image Segmentation Under Geometrical Priors
Vessel segmentation of retinal images is a key diagnostic capability in ophthalmology. This problem faces several challenges including low contrast, variable vessel size and thickness, and presence of interfering pathology such as micro-aneurysms and hemorrhages.
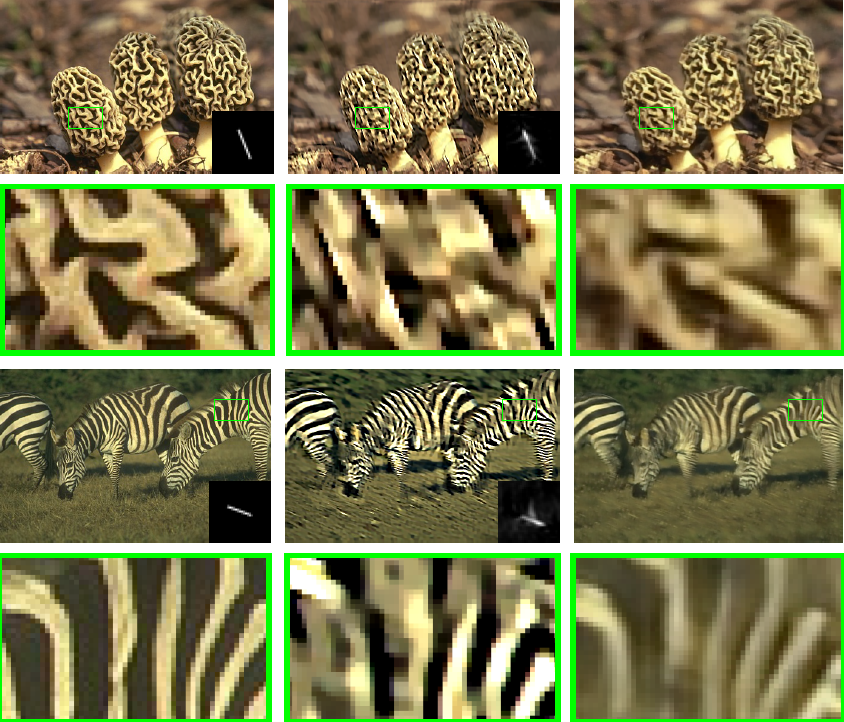
Efficient And Interpretable Deep Blind Image Deblurring Via Algorithm Unrolling
Blind image deblurring remains a topic of enduring interest. Learning based approaches, especially those that employ neural networks have emerged to complement traditional model based methods and in many cases achieve vastly enhanced performance. That said, neural network approaches are generally empirically designed and the underlying structures are difficult to interpret.
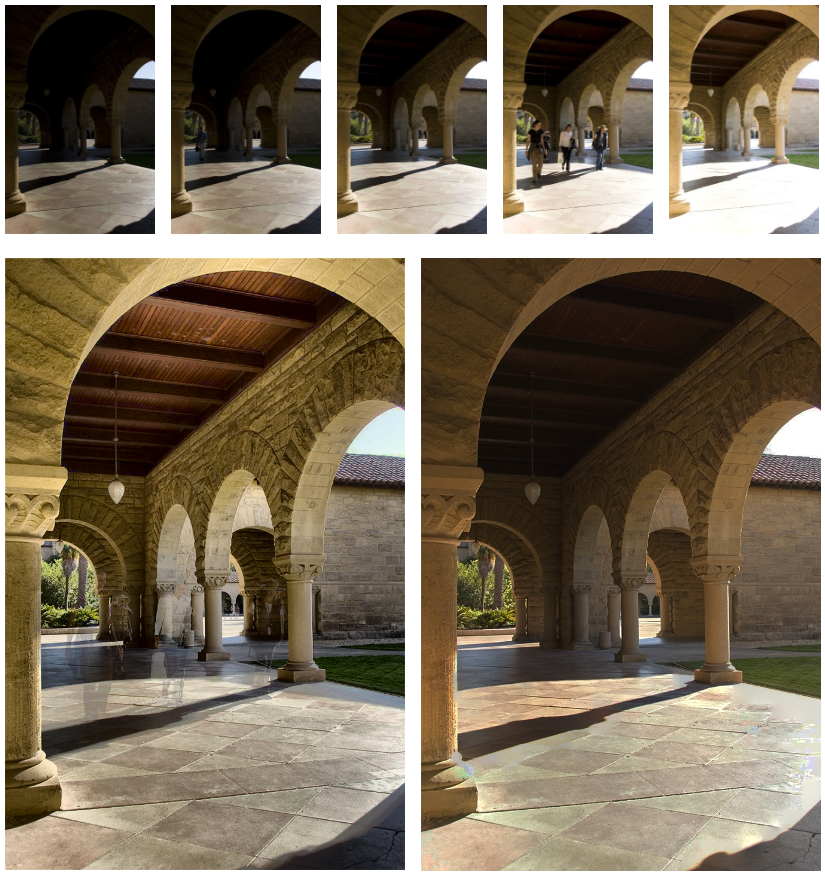
Ghost-Free High Dynamic Range Imaging
We propose a ghost-free high dynamic range (HDR) image synthesis algorithm using a low-rank matrix completion framework, which we call RM-HDR. Based on the assumption that irradiance maps are linearly related to low dynamic range (LDR) image exposures, we formulate ghost region detection as a rank minimization problem.
A MAP Estimation Framework for HDR Video Synthesis
High dynamic range (HDR) image synthesis from multiple low dynamic range (LDR) exposures continues to be a topic of great interest. Its extension to HDR video is also a topic of significant research interest due to its increasing demand and economic costs.
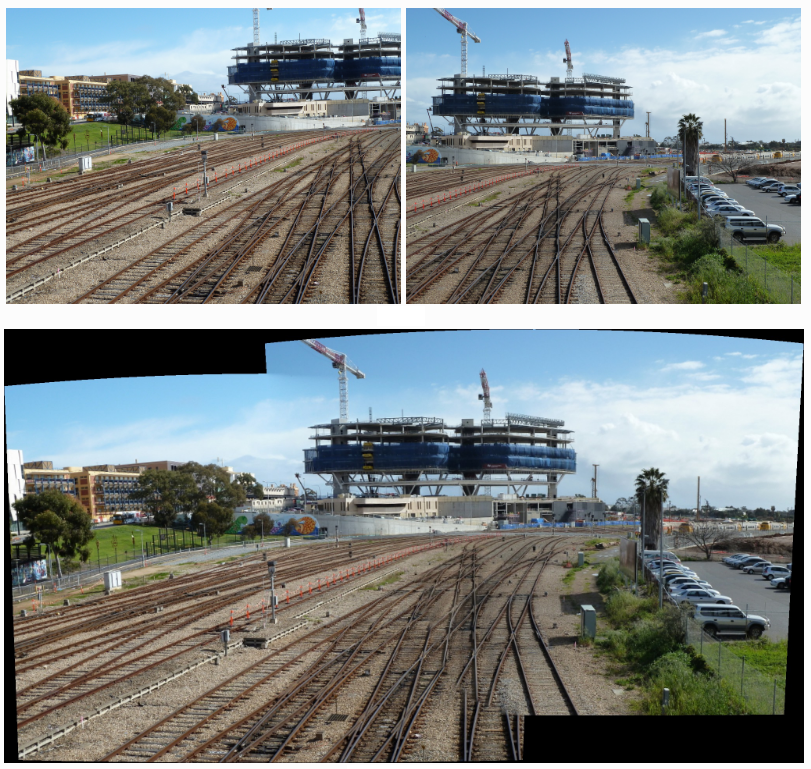
Bundle Robust Alignment for Panoramic Stitching
Here we study the problem of image alignment for panoramic stitching. Unlike most existing approaches that are feature-based, our algorithm works on pixels directly, and accounts for errors across the whole images globally.
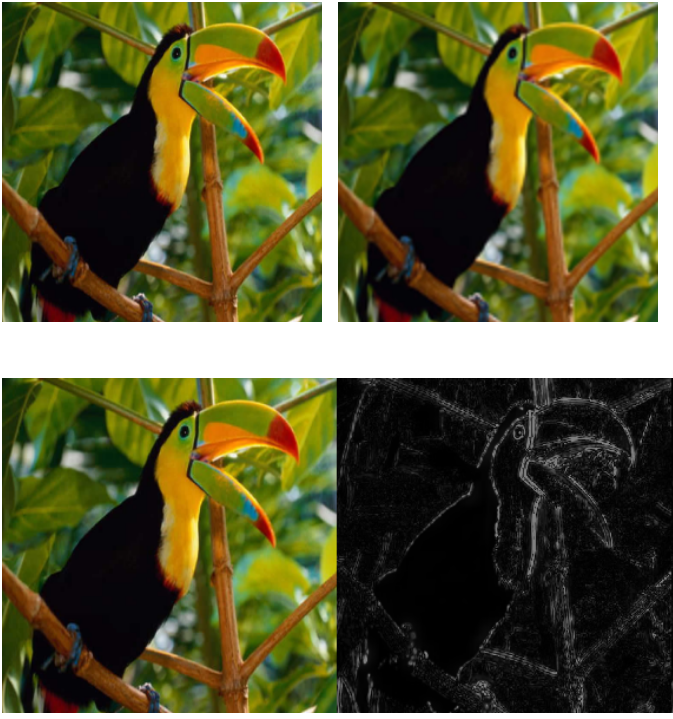
Sparsity-based Color Image Super Resolution via Exploiting Cross Channel Constraints
Sparsity constrained single image super-resolution (SR) has been of much recent interest. A typical approach involves sparsely representing patches in a low-resolution (LR) input image via a dictionary of example LR patches, and then using the coefficients of this representation to generate the highresolution (HR) output via an analogous HR dictionary.
Blind Image Deblurring Using Row-Column Sparse Representations
Blind image deblurring is a particularly challenging inverse problem where the blur kernel is unknown and must be estimated en route to recovering the de-blurred image. The problem is of strong practical relevance since many imaging devices such as cellphone cameras, must rely on deblurring algorithms to yield satisfactory image quality.